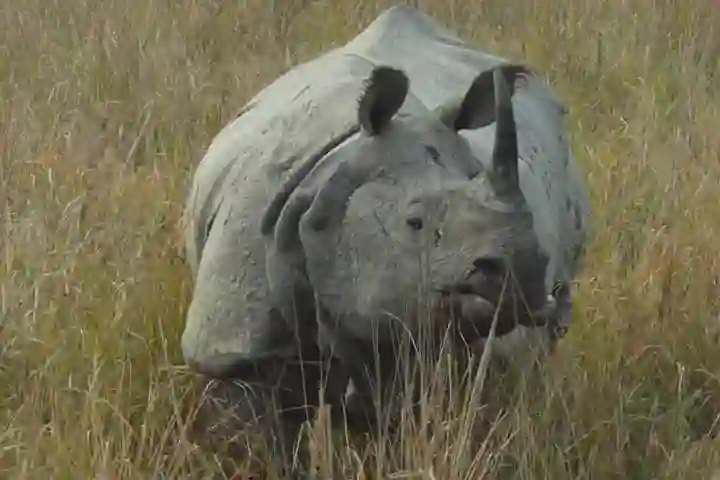
Scientists have always been intrigued about the relationships between the planet’s living rhinoceros species – a total of five in number. The answer to this question was difficult to get as most of the rhinos before the Pleistocene period became extinct.
An article in sciencedaily.com states that according to a report in the journal Cell, published on August 24, experts are now able to fill in the gaps in the revolutionary family tree of rhinos. This was possible by analysing genomes of five living species along with those of the three extinct species.
It was nearly 16 million years ago that the earliest split between the African and Eurasian branches took place as per the study. Also it was found that though the present rhino populations have lower genetic diversity and more cases of inbreeding than in the past, historically genetic diversity among them was always low.
Highlighting this, Love Dalen of the Centre for Palaeogenetics and the Swedish Museum of Natural History observed: “We can now show that the main branch in the rhinoceroses' tree of life is among geographic regions, Africa versus Eurasia, and not between the rhinos that have one versus two horns. The second important finding is that all rhinoceroses, even the extinct ones, have comparatively low genetic diversity. To some extent, this means that the low genetic diversity we see in present-day rhinos, which are all endangered, is partly a consequence of their biology.”
Also read: Rattlesnakes, masters of fatal deception
Airing his views on this issue, Mick Westbury of Denmark’s University of Copenhagen remarked: “All eight species generally displayed either a continual but slow decrease in population size over the last 2 million years, or continuously small population sizes over extended time periods. Continuously low population sizes may indicate that rhinoceroses in general are adapted to low levels of diversity."
The view seems to be in line with the absence of harmful mutations in the rhinos. Westbury felt that the creatures may have removed them in the last 10 decades, thereby enabling them to remain relatively healthy, notwithstanding the low genetic diversity.
The work on rhino saw the coming together of two groups of experts. Working separately on different rhino species were Dalen and Tom Glibert of University of Copenhagen. On becoming aware of this, they came together along with their other colleagues around the world, to do a comparative study. This study encompassed the living species as well as those which became extinct in the last Ice Age.
The joint study confronted certain challenges. One was that the genome data were different as it was based on ancient and modern DNA. To overcome this, a new analysis tool was developed.
Also read: Defying age, Cuttlefish has an "elephant's" memory
The findings as per Dalén are "partly good news, and partly not”. While the low genetic diversity is part of long history it has not caused any major health problems due to mutations and in-breeding.
Dalen added: "However, we also find that present-day rhinos have lower genetic diversity, and higher levels of inbreeding, compared to our historical and prehistoric rhinoceros genomes. This suggests that recent population declines caused by hunting and habitat destruction have had an impact on the genomes. This is not good, since low genetic diversity and high inbreeding may increase the risk of extinction in the present-day species."
The findings of the study will surely help in conservation and further studies of these endangered animals. Westbury remarked: "Now we know that the low diversity we see in contemporary individuals may not be indicative of an inability to recover, but instead a natural state of rhinoceros. We can better guide recovery programs to focus on increasing population size rather than individual genetic diversity."